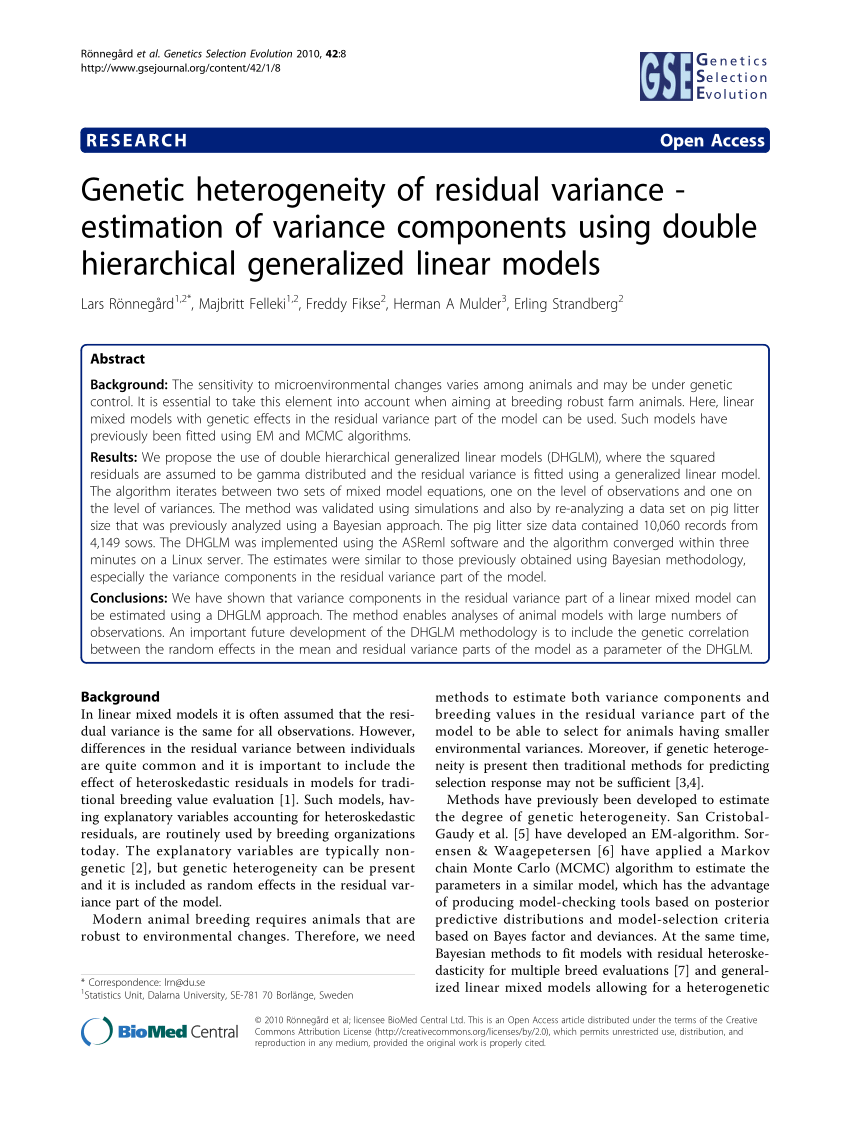

- VARIANCE COMPONENTS FOR FIXED EFFECTS ASREML CODE
- VARIANCE COMPONENTS FOR FIXED EFFECTS ASREML SERIES
WhichED = c ( "colId", "rowId", "fColRow", "colfRow", "surface" ), The corrected values of one time point are displayed in a table like the following: timeNumber In brief, separately for each measurement time \(t\), a spatial model is fitted for the trait \(y_t\), It will provide the user with either genotypic values or corrected values that can be used for further modeling. The aim of this document is to accurately separate the genetic effects from the spatial effects at each time point. It is also suitable for phenotyping platform data and has been tested on several datasets in the EPPN 2020 project. It has proven to be a good alternative to the classical AR1×AR1 modeling in the field (Velazco et al. It also provides the user with graphical outputs that are easy to interpret. This model corrects for spatial trends, row and column effects and has the advantage of avoiding the model selection step. An attractive alternative is the use of 2-dimensional P-spline surfaces, the SpATS model (Spatial Analysis of Trials using Splines, (Rodríguez-Álvarez et al.
VARIANCE COMPONENTS FOR FIXED EFFECTS ASREML SERIES
These models are sometimes difficult to fit and the selection of a best model is complicated, therefore preventing an automated phenotypic analysis of series of trials. Popular mixed models to separate spatial trends from treatment and genetic effects, rely on the use of autoregressive correlation functions defined on rows and columns (AR1×AR1) to model the local trends (Cullis, Smith, and Coombes 2006). In the same way as in field trials, platform experiments should obey standard principles for experimental design and statistical modeling. Taking into account these spatial trends is a prerequisite for precise estimation of genetic and treatment effects. For example, the spatial variability of incident light can go up to 100% between pots within a greenhouse (Cabrera-Bosquet et al. For the animal effect this proportion is (usually) equal to the heritability.Phenotyping facilities display spatial heterogeneity.
Conveniently however, WOMBAT estimates not only the absolute variance components, but also the proportion of the phenotypic variation accounted for by the different random effects, along with their sampling error. We can calculate the heritability by hand from the variance component estimates provided in SumEstimates.out. Posterior.mode(posterior.heritability1.1) HPDinterval(posterior.heritability1.1,0.95) Summary(model)$varcomp/sum(summary(model)$varcomp)
VARIANCE COMPONENTS FOR FIXED EFFECTS ASREML CODE
If additive genetic variance is the 1st random effect, the following code will work - otherwise change the number 1 for the appropriate row. By extracting variance components from summary(model)$varcomp. By hand - this is easiest if you are moving output into another program which can do calculations, Excel for example. There are two ways to calculate heritability in ASReml-R. pin file accordingly or you will get the wrong answer! NOTE - if you change the random effects stucture of your model in. H h2 1 3 #divides 1 (VA) by 3 (VP) to calculate h2 pin file to calculate heritability from these components migt beį VP 1+2 #adds components 1 and 2 to make a 3rd variance denoted VP additive genetic component), the second will be the residual variance. The primary output file (.asr) will contain two variance components. pin) is used to caculate functions of estimated variance components ad their associated standard errors. In ASReml a second command file (with extension. So in the simplest case in which we just have an additive genetic (V A) and a residual variance component (V R), h 2 = V A / V A+V R. Divide additive genetic variance by the total phenotypic variance. For further details see Wilson et al 2008 (pdf): Why h 2 does not always equal V A / V P?Īdd up all variance components to find the total phenotypic variance (once fixed effects have been corrected for). As variance components are calulated after fixed effects have been accounted for, the addition of fixed effects into the model can cause the estimate of h 2 to change. Heritability (or h 2) measures the proportion of phenotypic variance accounted for by additive genetic effects and is a central property of quantitative genetics.
